8 NED with open LLMs and domain ontologies
This chapter introduces a practical, domain-agnostic approach to Named Entity Disambiguation that marries open large language models with rich domain ontologies. It begins by motivating the need to go beyond traditional tools such as ScispaCy, which, while strong in biomedical contexts, can be hard to update, are tied to a single domain, and underuse the relational knowledge embedded in ontologies. The proposed solution runs locally on consumer hardware using an open model (Llama 3.1 8B via Ollama) and leverages SNOMED as a reference ontology, addressing privacy, latency, and extensibility while explicitly exploiting ontology structure during disambiguation.
The implementation loads SNOMED into Neo4j, normalizing many relationship types under a single relationship with a type property and introducing a dedicated hierarchical link to propagate semantic categories from top-level concepts down the taxonomy. These propagated categories feed an ontology-guided NER step in which the LLM is prompted to extract only ontology-derived entity types; character offsets are then computed deterministically in post-processing. Candidate selection does not rely on the LLM: it uses Neo4j full-text search constrained by the NER labels to retrieve plausible SNOMED candidates for each mention (e.g., multiple interpretations for “Zika”), establishing a focused pool for the final decision.
Candidate disambiguation combines graph algorithms and LLM reasoning in three steps: shortest-path detection between candidates within the ontology (with hub filtering and limited hops for relevance), path-to-text translation that turns the discovered graph paths into natural-language statements, and summarization that condenses many sentences into a compact context. The final LLM prompt uses this distilled, ontology-grounded context to pick the best candidate per mention—e.g., preferring “Congenital Zika virus infection” when “microcephaly” co-occurs. The result is an end-to-end NED workflow that is accurate in sparse or ambiguous contexts, portable across domains with suitable ontologies, privacy-friendly when run locally, and extensible with improvements such as vector-based candidate retrieval.
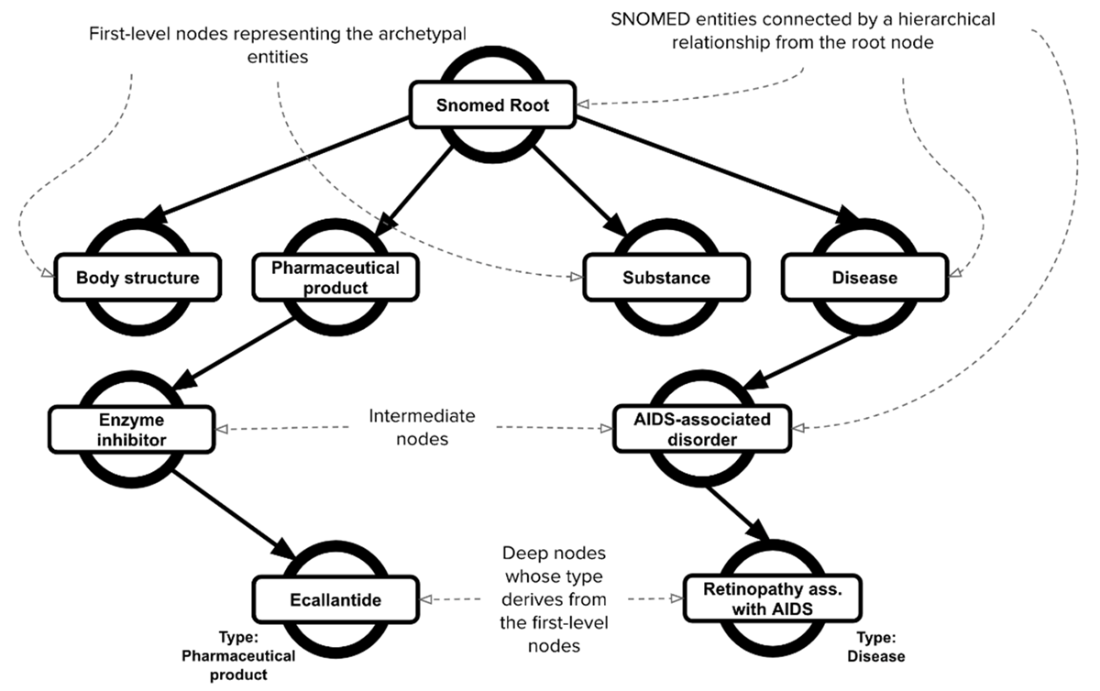
A sample of the hierarchical structure of the SNOMED ontology. Leveraging this hierarchical structure, nodes located on a deeper level, such as “Ecallantide” and “Retinopathy associated with AIDS”, can be categorized using the information from the first-level nodes, such as “Pharmaceutical product” and “Disease”, which represent the archetypal entities of the ontology.
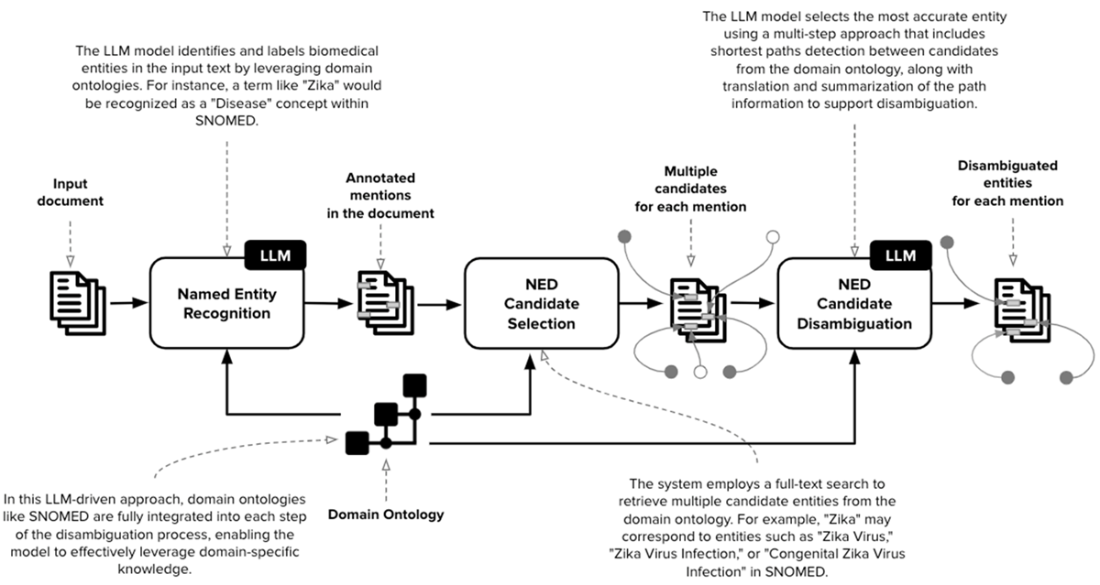
A comprehensive workflow for a Named Entity Disambiguation (NED) system designed to leverage Large Language Models (LLMs) and domain-specific ontologies, such as SNOMED, for biomedical text processing. This workflow is organized into distinct stages, each involving various processes and interactions between the LLM and domain ontology to disambiguate entities within input text accurately.
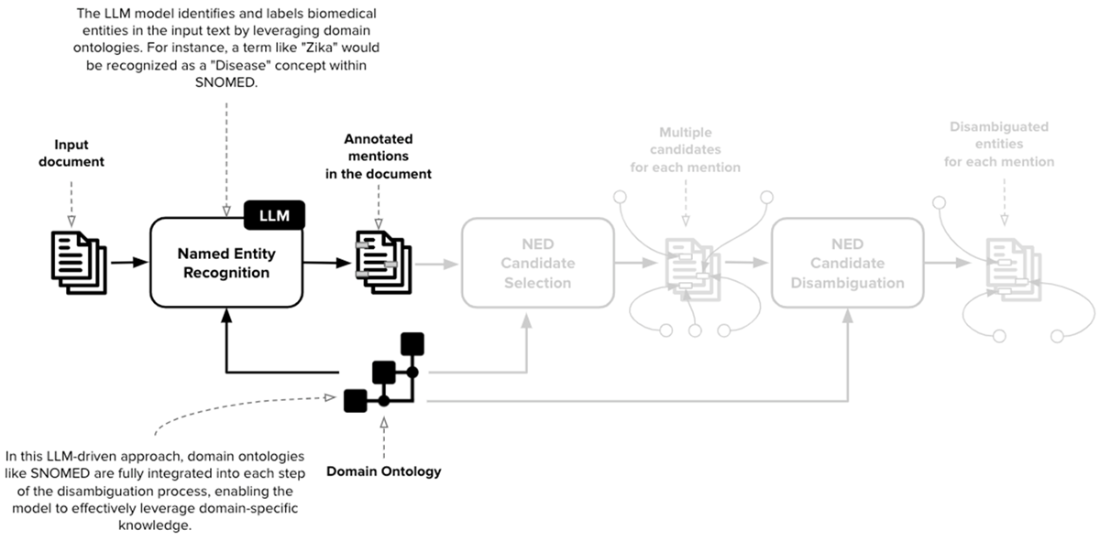
The first stage of NED is Named Entity Recognition (NER). In our scenario, the collection of named entities is derived directly from the ontology. In SNOMED, the categories are defined by the first-level nodes of the ontology, whose information is propagated to all the other nodes.
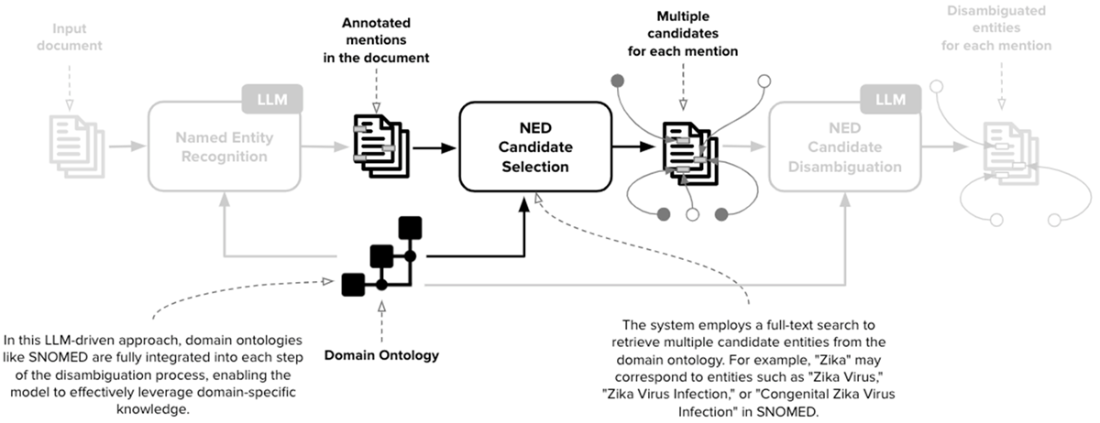
The second stage of NED is the Candidate Selection. In this scenario, for each entity mention detected in the previous step, this stage retrieves a collection of potential candidates that can refer to each mention. The current implementation employs a full-text search but can be potentially extended with more advanced techniques.
The third stage of NED is the Candidate Disambiguation. The key goal is to select the best match among all the possible candidates identified in the previous step. To reach this goal, the disambiguation phase exploits an advanced approach combining graph-based algorithms (shortest path detection) and LLMs.
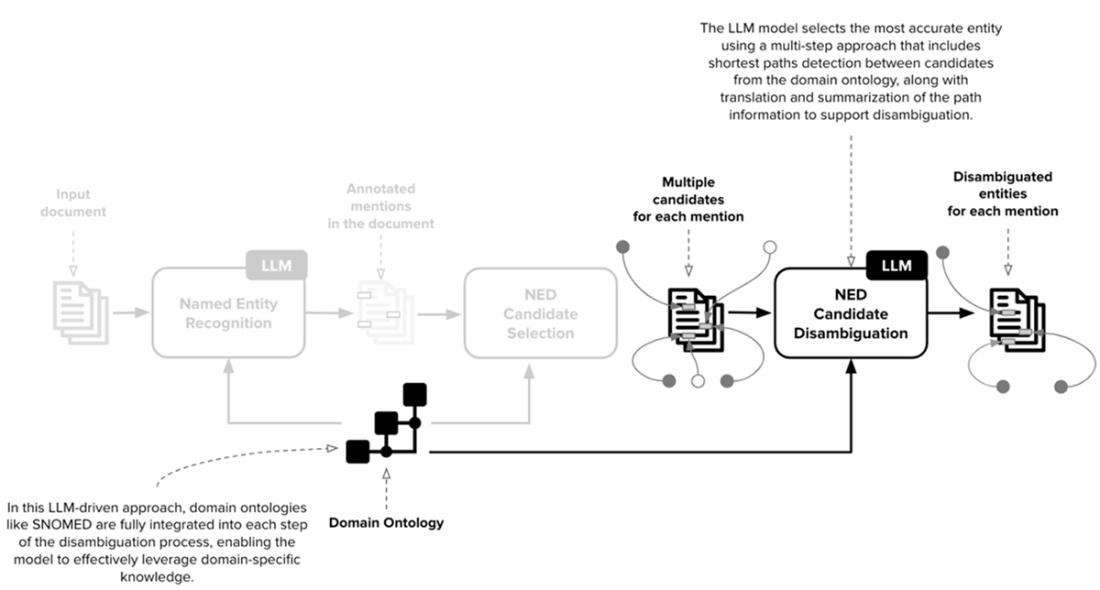
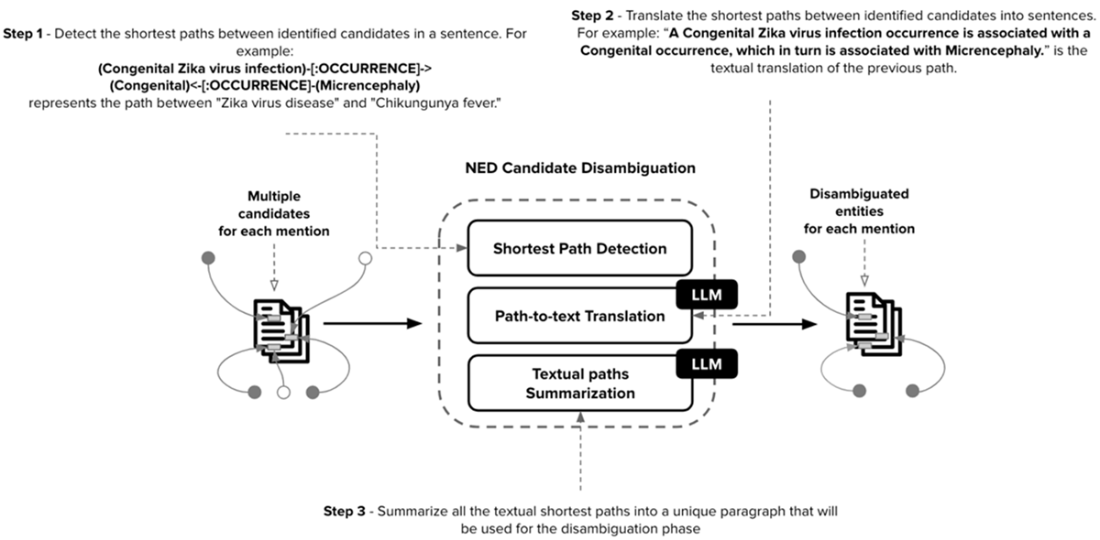
NED Candidate Disambiguation is divided into three main phases: (1) Shortest Path Detection between all the candidates related to the different entity mentions in the sentence. (2) Path-to-text Translation to transform detected paths into natural language sentences. (3) Textual paths summarization to summarize all the natural language sentences into a unique and valuable piece of text useful for the disambiguation
Summary
- Named Entity Disambiguation (NED) is essential for accurately identifying and distinguishing entities in complex domains, particularly in biomedical text.
- Traditional Natural Language Processing (NLP) tools such as ScispaCy have some limitations:
- they can not be used in diverse domains, they can not leverage the relationships between entities, and their reference knowledge can not be extended and updated.
- The combination of general-purpose Large Language Models (LLMs) and domain ontologies allows us to address these issues: LLMs can be driven by the continuously updated knowledge incorporated by the ontology and leverage its relational structure.
- To reach this goal, we can deploy a flexible end-to-end process for NED, including multiple phases involving LLMs and domain ontologies, such as Named Entity Recognition (NER), NED Candidate Selection, and NED Candidate Disambiguation.
- To fully leverage the capabilities of LLMs combined with the graph dimension of domain ontologies, the disambiguation phase is divided into three different stages:
- Shortest-path detection.
- Path-to-text translation.
- Textual path summarization.
- Future NED applications can leverage this framework and adapt to different domains, which are characterized by rich ontologies describing the relational nature of their specific entities.
FAQ
What limitations of traditional NED tools like ScispaCy does this chapter address?
ScispaCy is domain-specific (biomedical), hard to expand/update with new entities and aliases, and it underutilizes knowledge base structure by not leveraging relationships and paths. The chapter addresses these gaps by combining open LLMs with domain ontologies and graph algorithms to use hierarchical and relational context during disambiguation.Why combine open LLMs with domain ontologies for NED?
LLMs provide broad linguistic competence, while ontologies like SNOMED contribute precise, curated structure. Together they: 1) generalize across domains; 2) ground decisions in canonical concepts; 3) exploit hierarchical/relational context; 4) improve accuracy under sparse local context; and 5) can run locally for privacy with tools like Ollama.What is SNOMED, and which files are ingested?
SNOMED CT is a comprehensive, multilingual clinical terminology with 450k+ concepts and rich relationships. The chapter ingests two TSV files: 1) sct2_Description_Full-en_*.txt for names and aliases; 2) sct2_Relationship_Full_*.txt for relationships (triples and metadata).How is SNOMED modeled and loaded in Neo4j?
- Nodes: SnomedEntity(id, name, aliases)- Relationships: a generic SNOMED_RELATION with type and id; plus a dedicated SNOMED_IS_A for hierarchy
- Indexes/constraints: unique id, indexes for name, relation id/type, and a full-text name index
- Names/aliases are added to both nodes and relationships from the description file.
How are NER categories derived from SNOMED?
First-level ontology nodes (e.g., Disease, Organism, Substance, Event) are propagated down the SNOMED_IS_A hierarchy using APOC traversal. The collected types (n.type) become the allowed labels in the NER prompt, ensuring entity extraction aligns with ontology categories.How does candidate selection work, and why isn’t the LLM used here?
Candidate selection uses Neo4j full-text search on SnomedEntity names/aliases with fuzzy matching, filtered by the NER label to limit scope. It returns (snomed_id, name) candidates per mention. LLMs aren’t used because the ontology is too large for prompts and we want exact retrieval from the ground-truth KB rather than model priors.What is the shortest-path strategy for disambiguation?
For candidates across co-occurring mentions in a sentence: 1) compute allShortestPaths (1–2 hops) between candidate pairs; 2) exclude high-degree “hub” nodes via GDS degree to avoid generic links; 3) format paths with relationship directions and types; these paths become evidence used downstream in LLM prompts.How are graph paths turned into text, and why summarize them?
Paths like (A)-[:REL]->(B)<-[:REL]-(C) are translated by an LLM into clear sentences that preserve exact entity names. A second prompt summarizes multiple sentences into a concise “context” to reduce token load and highlight salient relationships that guide the final disambiguation.How do I run Llama 3.1 8B locally with Ollama and call it from Python?
- Install and run: “ollama serve”, then “ollama pull llama3.1:latest”- Use OpenAI-compatible Chat Completions via base_url http://localhost:11434/v1 and model "llama3.1:latest"
- Set temperature low (e.g., 0) for deterministic NED outputs.
What enhancements and practical tips does the chapter suggest?
- Add vector search to boost candidate recall alongside full-text- Tune hub filtering thresholds and path lengths; cache computed paths
- Post-process character offsets deterministically to fix LLM span errors
- Extend to other domains by swapping in rich ontologies
- Evaluate with standard NED metrics; incrementally update aliases and entities; leverage multilingual capabilities of Llama 3.1 where helpful.
 Knowledge Graphs and LLMs in Action ebook for free
Knowledge Graphs and LLMs in Action ebook for free